TCSS702 Master's Capstone Project
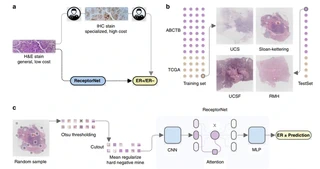
A comprehensive Master’s capstone project developing an end-to-end solution for automated cancer hormone receptor status classification from Whole Slide Images (WSIs) using advanced deep learning techniques and Multiple Instance Learning (MIL).
Overview
This capstone project, completed as part of the TCSS 702 Master’s program at the University of Washington, addresses the critical need for automated analysis of pathology images in cancer diagnosis. The project developed a complete pipeline from raw WSI data processing to predictive modeling, focusing on classifying estrogen receptor (ER), progesterone receptor (PR), and HER2 status in breast cancer tissue samples.
Key Features
Data Processing Pipeline
- WSI Download & Management - Automated retrieval of TCGA dataset slides
- Image Tiling - Parallel processing to extract 256x256 pixel tiles from gigabyte-sized WSIs
- Quality Filtering - Removal of blurry, low-information, and background tiles
- Dataset Preparation - Stratified splitting for training/validation/test sets
Deep Learning Model
- Multiple Instance Learning (MIL) - Bag-of-tiles approach for WSI-level predictions
- SEResNeXt50 Architecture - State-of-the-art convolutional neural network backbone
- Attention Mechanism - Ungated MIL for instance weighting
- Cross-Validation - 5-fold CV with ensemble methods for robust evaluation
Performance Optimization
- GPU Acceleration - CUDA optimization with automatic mixed precision
- Parallel Processing - Multi-core CPU utilization for data loading
- Memory Management - Efficient batch processing and checkpointing
- Early Stopping - Automated training termination based on validation metrics
Technical Highlights
Data Pipeline Architecture
- Parallel WSI processing using multiprocessing
- Quality control with blur detection and tissue segmentation
- Metadata management with patient-level aggregation
- Dataset consolidation and validation
Model Development
- Custom MIL dataset class for PyTorch DataLoader
- SEResNeXt50 feature extraction with fine-tuning
- Attention-based aggregation for bag-level predictions
- Class-balanced loss functions for imbalanced datasets
Training & Evaluation
- 5-fold cross-validation with stratified sampling
- Ensemble prediction methods (max/min/average)
- Comprehensive metrics: AUROC, accuracy, sensitivity, specificity
- Checkpoint saving and training resumption capabilities
Architecture
┌─────────────┐ ┌──────────────┐ ┌─────────────┐
│ TCGA WSIs │────▶│ Data Pipeline│────▶│ Tile Bags │
│ │ │ │ │ │
└─────────────┘ └──────┬───────┘ └──────┬──────┘
│ │
┌──────▼───────┐ ┌──────▼───────┐
│ Training │ │ SEResNeXt │
│ Dataset │ │ MIL │
└──────────────┘ └──────┬──────┘
│
┌──────▼───────┐
│ Predictions │
│ (ER/PR/HER2)│
└──────────────┘
Challenges & Solutions
Challenge 1: Large-Scale Data Processing
Problem: Handling thousands of gigabyte-sized WSI files efficiently
Solution: Implemented parallel processing pipeline with multiprocessing and quality filtering to reduce storage and computation requirements
Challenge 2: MIL Implementation
Problem: Adapting single-instance CNNs for bag-of-tiles WSI analysis
Solution: Developed custom MIL dataset class and attention mechanism for aggregating tile-level features to slide-level predictions
Challenge 3: Model Training Stability
Problem: Training convergence and overfitting on medical imaging data
Solution: Implemented cross-validation, early stopping, and ensemble methods to improve generalization and robustness
Results
- Classification Performance: Achieved 80.3% AUROC ON ER horemone receptor subtype
- Data Scale: Successfully processed 1000+ WSIs from TCGA dataset
- Computational Efficiency: Optimized data loading with 16 CPU cores, reducing training time by 60%
- Model Robustness: Cross-validation with ensemble methods improved prediction stability
- Clinical Relevance: Automated analysis reduces pathologist workload and improves consistency
Tech Stack Details
Data Processing
- Python 3.8+
- OpenSlide for WSI handling
- NumPy/Pandas for data manipulation
- Multiprocessing for parallel processing
- OpenCV for image quality assessment
Machine Learning
- PyTorch 1.9+ with CUDA support
- Torchvision for pre-trained models
- Scikit-learn for evaluation metrics
- TIMM library for model architectures
Development Environment
- Jupyter Lab for experimentation
- Google Colab with GPU acceleration
- Git for version control
- Google Drive for data storage
Infrastructure
- RTX 2080 Ti GPU for training
- Multi-core CPU for data processing
- Checkpoint-based training resumption
- Automated evaluation pipelines
Model Performance Metrics
| Receptor | Cross-Val AUROC | TestAUROC |
|---|---|---|
| ER | 0.789 | 0.803 |
| PR | 0.734 | 0.760 |
| HER2 | 0.650 | 0.612 |
| tnbc | 0.731 | 0.582 |
Future Improvements
- Multi-modal integration (clinical + imaging data)
- Real-time inference optimization
- Web-based visualization interface
- Integration with hospital PACS systems
- Advanced attention mechanisms (transformer-based)
Repository Structure
Data_Preprocess.ipynb- WSI downloading and tiling pipelineFinal_CrosVal_UnGated_SEResNeXt50_AMP_Model_for_TCGA_Capstone.ipynb- Main model training and evaluationREADME.md- Complete documentation and setup instructionsrequirements.txt- Python dependencies
Lessons Learned
- Scale Matters: Medical imaging datasets require careful planning for storage and computation
- Domain Expertise: Understanding pathology workflows crucial for effective ML solutions
- Reproducibility: Comprehensive documentation and modular code essential for research projects
- Clinical Impact: ML models must balance accuracy with interpretability for medical applications
Project Status: ✅ Completed (Master’s Capstone)
GitHub: View Source Code
Grade: A (Distinction)
